Evolution of Modularity
post by johnswentworth · 2019-11-14T06:49:04.112Z · LW · GW · 12 commentsContents
12 comments
This post is based on chapter 15 of Uri Alon’s book An Introduction to Systems Biology: Design Principles of Biological Circuits. See the book for more details and citations; see here [LW · GW] for a review of most of the rest of the book.
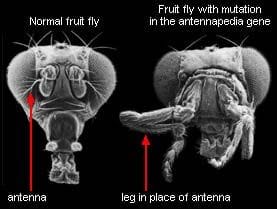
Fun fact: biological systems are highly modular, at multiple different scales. This can be quantified and verified statistically, e.g. by mapping out protein networks and algorithmically partitioning them into parts, then comparing the connectivity of the parts. It can also be seen more qualitatively in everyday biological work: proteins have subunits which retain their function when fused to other proteins, receptor circuits can be swapped out to make bacteria follow different chemical gradients, manipulating specific genes can turn a fly’s antennae into legs, organs perform specific functions, etc, etc.
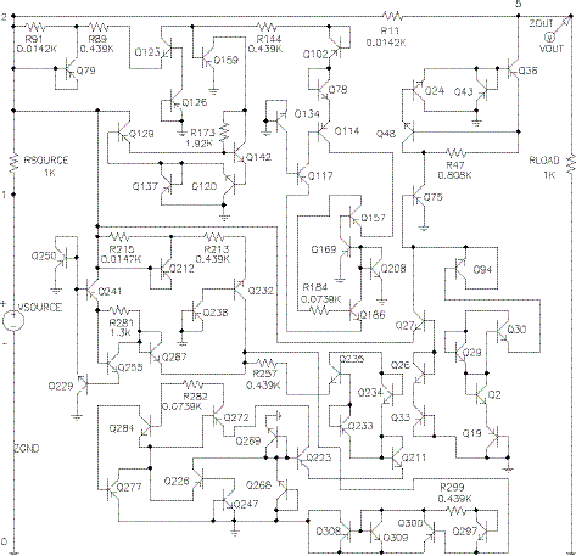
On the other hand, systems designed by genetic algorithms (aka simulated evolution) are decidedly not modular. This can also be quantified and verified statistically. Qualitatively, examining the outputs of genetic algorithms confirms the statistics: they’re a mess.
So: what is the difference between real-world biological evolution vs typical genetic algorithms, which leads one to produce modular designs and the other to produce non-modular designs?
Kashtan & Alon tackle the problem by evolving logic circuits under various conditions. They confirm that simply optimizing the circuit to compute a particular function, with random inputs used for selection, results in highly non-modular circuits. However, they are able to obtain modular circuits using “modularly varying goals” (MVG).
The idea is to change the reward function every so often (the authors switch it out every 20 generations). Of course, if we just use completely random reward functions, then evolution doesn’t learn anything. Instead, we use “modularly varying” goal functions: we only swap one or two little pieces in the (modular) objective function. An example from the book:
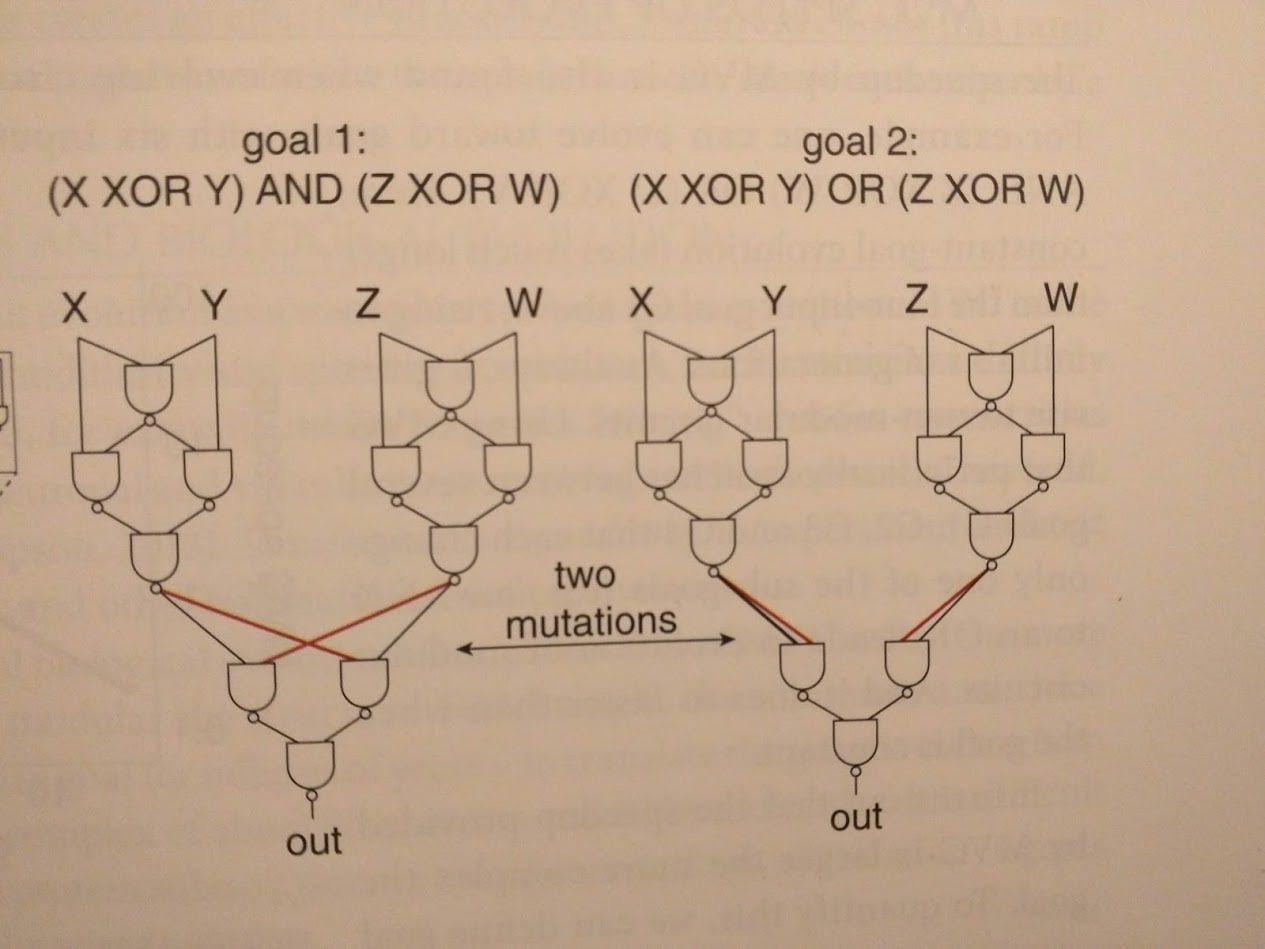
The upshot is that our different goal functions generally use similar sub-functions - suggesting that they share sub-goals for evolution to learn. Sure enough, circuits evolved using MVG have modular structure, reflecting the modular structure of the goals.
(Interestingly, MVG also dramatically accelerates evolution - circuits reach a given performance level much faster under MVG than under a fixed goal, despite needing to change behavior every 20 generations. See either the book or the paper for more on that.)
How realistic is MVG as a model for biological evolution? I haven’t seen quantitative evidence, but qualitative evidence is easy to spot. MVG as a theory of biological modularity predicts that highly variable subgoals will result in modular structure, whereas static subgoals will result in a non-modular mess. Alon’s book gives several examples:
- Chemotaxis: different bacteria need to pursue/avoid different chemicals, with different computational needs and different speed/energy trade-offs, in various combinations. The result is modularity: separate components for sensing, processing and motion.
- Animals need to breathe, eat, move, and reproduce. A new environment might have different food or require different motions, independent of respiration or reproduction - or vice versa. Since these requirements vary more-or-less independently in the environment, animals evolve modular systems to deal with them: digestive tract, lungs, etc.
- Ribosomes, as an anti-example: the functional requirements of a ribosome hardly vary at all, so they end up non-modular. They have pieces, but most pieces do not have an obvious distinct function.
To sum it up: modularity in the system evolves to match modularity in the environment.
12 comments
Comments sorted by top scores.
comment by Kaj_Sotala · 2019-11-14T09:39:05.598Z · LW(p) · GW(p)
There is also the suggestion that having connection costs imposes modularity:
We investigate an alternate hypothesis [than the MVG one] that has been suggested, but heretofore untested, which is that modularity evolves not because it conveys evolvability, but as a byproduct from selection to reduce connection costs in a network (figure 1) [9,16]. Such costs include manufacturing connections, maintaining them, the energy to transmit along them and signal delays, all of which increase as a function of con- nection length and number [9,17 –19]. The concept of connection costs is straightforward in networks with physical connections (e.g. neural networks), but costs and physical limits on the number of possible connections may also tend to limit interactions in other types of networks such as genetic and metabolic pathways. For example, adding more connections in a signalling pathway might delay the time that it takes to output a critical response; adding regulation of a gene via more transcription factors may be difficult or impossible after a certain number of proximal DNA binding sites are occupied, and increases the time and material required for genome replication and regulation; and adding more protein–protein interactions to a system may become increasingly difficult as more of the remaining surface area is taken up by other binding interactions. Future work is needed to investigate these and other hypotheses regarding costs in cellular networks. The strongest evidence that biological networks face direct selection to minimize connection costs comes from the vascular system [20] and from nervous systems, including the brain, where multiple studies suggest that the summed length of the wiring diagram has been minimized, either by reducing long connections or by optimizing the placement of neurons [9,17 –19,21 –23]. Founding [16] and modern [9] neuroscientists have hypothesized that direct selection to minimize connection costs may, as a side-effect, cause modularity. [...]
Given the impracticality of observing modularity evolve in biological systems, we follow most research on the subject by conducting experiments in computational systems with evolutionary dynamics [4,11,13]. Specifically, we use a well- studied system from the MVG investigations [13,14,27]: evolving networks to solve pattern-recognition tasks and Boolean logic tasks (§4). [...]
Replies from: johnswentworthAfter 25 000 generations in an unchanging environment (L-AND-R), treatments selected to maximize performance and minimize connection costs (P&CC) produce significantly more modular networks than treatments maximizing per- formance alone (PA)
↑ comment by johnswentworth · 2019-11-14T18:13:42.363Z · LW(p) · GW(p)
Yeah, Alon briefly mentions that line of study as well, although he doesn't discuss it much. Personally, I think connection costs are less likely to be the main driver of biological modularity in general, for two main reasons:
- If connection costs were a taut constraint, then we'd expect to see connection costs taking up a large fraction of the organism's resources. I don't think that's true for most organisms most of the time (though the human brain is arguably an exception). And qualitatively, if we look at the cost of e.g. signalling molecules in a bacteria, they're just not that expensive - mainly because they don't need very high copy number.
- Connection costs are not a robust way to produce modularity - we need a delicate balance between cost and benefit, so that neither overwhelms the other. Given how universal modularity is in biology, across so many levels of organization and basically all known organisms, it seems like a less delicate mechanism is needed to explain it.
I do find it plausible that connection cost is a major driver in some specific systems - in particular, the sanity checks pass for the human brain. But I doubt that it's the main cause of modularity across so many different systems in biology.
comment by johnswentworth · 2021-01-10T19:15:01.211Z · LW(p) · GW(p)
The material here is one seed of a worldview which I've updated toward a lot more over the past year. Some other posts which involve the theme include Science in a High Dimensional World [LW · GW], What is Abstraction? [LW · GW], Alignment by Default [LW · GW], and the companion post to this one Book Review: Design Principles of Biological Circuits [LW · GW].
Two ideas unify all of these:
- Our universe has a simplifying structure: it abstracts well, implying a particular kind of modularity.
- Goal-oriented systems in our universe tend to evolve a modular structure which reflects the structure of the universe.
One major corollary of these two ideas is that goal-oriented systems will tend to evolve similar modular structures, reflecting the relevant parts of their environment. Systems to which this applies include organisms, machine learning algorithms, and the learning performed by the human brain. In particular, this suggests that biological systems and trained deep learning systems are likely to have modular, human-interpretable internal structure. (At least, interpretable by humans familiar with the environment in which the organism/ML system evolved.)
This post talks about some of the evidence behind this model: biological systems are indeed quite modular, and simulated evolution experiments find that circuits do indeed evolve modular structure reflecting the modular structure of environmental variations. The companion post [LW · GW] reviews the rest of the book, which makes the case that the internals of biological systems are indeed quite interpretable.
On the deep learning side, researchers also find considerable modularity in trained neural nets, and direct examination of internal structures reveals plenty of human-recognizable features.
Going forward, this view is in need of a more formal and general model, ideally one which would let us empirically test key predictions - e.g. check the extent to which different systems learn similar features, or whether learned features in neural nets satisfy the expected abstraction conditions, as well as tell us how to look for environment-reflecting structures in evolved/trained systems.
comment by Vanessa Kosoy (vanessa-kosoy) · 2020-12-03T11:52:47.522Z · LW(p) · GW(p)
I liked this post for talking about how evolution produces modularity (contrary to what is often said in this community!). This is something I suspected myself but it's nice to see it explained clearly, with backing evidence.
comment by habryka (habryka4) · 2020-12-15T06:36:13.468Z · LW(p) · GW(p)
Coming back to this post, I have some thoughts related to it that connect this more directly to AI Alignment that I want to write up, and that I think make this post more important than I initially thought. Hence nominating it for the review.
Replies from: johnswentworth↑ comment by johnswentworth · 2020-12-24T15:42:28.115Z · LW(p) · GW(p)
I'm curious to hear these thoughts.
comment by Thomas Kwa (thomas-kwa) · 2022-04-04T17:44:53.509Z · LW(p) · GW(p)
Maybe the hypothesis that environmental modularity drives agent modularity can be tested empirically using something like Deepmind's XLand. Fix the size of the space of tasks in the training distribution. Then find some way to define environmental modularity of a subset of tasks. Then test: do RL agents trained on a modular space of tasks have a more modular structure than RL agents trained on an arbitrary space of tasks?
comment by Kenoubi · 2022-02-24T14:16:03.149Z · LW(p) · GW(p)
Aren't "modularly varying reward functions" exactly what D𝜋's Self-Organised Neural Networks [LW · GW] accomplish? Each example in the training data is a module of the reward function. By only learning on the training examples that are currently hardest for the network, we make those examples easier and thus implicitly swap them out of the "examples currently hardest for the network" set.
comment by tailcalled · 2022-01-12T09:24:29.229Z · LW(p) · GW(p)
There's another form of modular variation with e.g. eating than just modularly varying foods in the environment, namely: we don't eat all the time. Instead we have some periods where we eat and other periods where we do other stuff. I wonder if this also contributes to modularity, e.g. it makes it necessary for it to be possible to "activate" and "deactivate" an eating mode while the rest of the body just does whatever.